We need to talk…
…said Pramesh, the Artificial Intelligence researcher.
‘I agree. I’m curious. The genetics of brain wiring is not really part of the AI discussion, is it?’ said Minda, the developmental geneticist.
![]() Pramesh: ‘That’s true. For decades AI tried to have nothing to do with biological messiness. Yet, just in the last few years, AI has become synonymous with neural networks and deep learning.’
Pramesh: ‘That’s true. For decades AI tried to have nothing to do with biological messiness. Yet, just in the last few years, AI has become synonymous with neural networks and deep learning.’
![]() Minda: ‘Well, I study genes and the development of neural networks. Why are there no genes and no development in your neural networks?’
Minda: ‘Well, I study genes and the development of neural networks. Why are there no genes and no development in your neural networks?’
![]() We design the networks. They become smart through learning.
We design the networks. They become smart through learning.
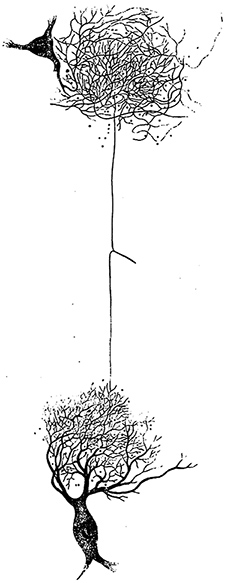
![]() Genetically encoded development can wire up a brain before it learns, a spider can build a web, and a fly can fly. They are already smart when they are born.
Genetically encoded development can wire up a brain before it learns, a spider can build a web, and a fly can fly. They are already smart when they are born.
![]() Artificial neural networks are typically designed with an initially random connectivity and then turned on to learn, so they need no prior development. The individual connection strengths change through big data or self-learning. Self-learning has been particularly successful recently – and isn’t self-learning what babies do?
Artificial neural networks are typically designed with an initially random connectivity and then turned on to learn, so they need no prior development. The individual connection strengths change through big data or self-learning. Self-learning has been particularly successful recently – and isn’t self-learning what babies do?
![]() Babies are not born with a randomly connected brain and then turned on to learn. But, funny enough, 100 years ago many scientists believed it would have to be like that. They were just as baffled as we are today about the amount of information needed to specify connectivity in the brain; they thought it must to come from learning. Kinda the same idea: a more or less randomly connected brain before you turn it on, and then learning makes it smart. This all changed when we started to study and understand developmental genetics – but we are talking more than 60 years ago! I find it hard to imagine that your artificial neural networks are stuck in that age.
Babies are not born with a randomly connected brain and then turned on to learn. But, funny enough, 100 years ago many scientists believed it would have to be like that. They were just as baffled as we are today about the amount of information needed to specify connectivity in the brain; they thought it must to come from learning. Kinda the same idea: a more or less randomly connected brain before you turn it on, and then learning makes it smart. This all changed when we started to study and understand developmental genetics – but we are talking more than 60 years ago! I find it hard to imagine that your artificial neural networks are stuck in that age.
![]() Well, do you understand how the genes encode connections in the brain and could you tell me how to implement that? I have sat through lecture after lecture on this and still feel I have learned little more than details about molecules. And every time somebody looks at the same molecule in a different context, things seem to be different. How does that help?
Well, do you understand how the genes encode connections in the brain and could you tell me how to implement that? I have sat through lecture after lecture on this and still feel I have learned little more than details about molecules. And every time somebody looks at the same molecule in a different context, things seem to be different. How does that help?
![]() We know molecules that tell neurons where to grow, and where to make synaptic
We know molecules that tell neurons where to grow, and where to make synaptic
connections with other neurons. But there is a lot to study, and we are finding more and more examples along these lines. There’s a general principle here.
![]() This is exactly where I am lost. What’s the principle and what’s the endpoint of this? How can I use this information to improve AI?
This is exactly where I am lost. What’s the principle and what’s the endpoint of this? How can I use this information to improve AI?
![]() I would say, theoretically, if you knew the entire molecular code for connectivity as it develops, you’d have a complete plan for brain development. We may not know all the molecules yet, but they are all encoded by just a few thousand genes; there is no fundamental enigma as to how it works.
I would say, theoretically, if you knew the entire molecular code for connectivity as it develops, you’d have a complete plan for brain development. We may not know all the molecules yet, but they are all encoded by just a few thousand genes; there is no fundamental enigma as to how it works.
![]() But the genetic code contains so much less information than the connectivity of the brain. Where do you think this information is coming from?
But the genetic code contains so much less information than the connectivity of the brain. Where do you think this information is coming from?
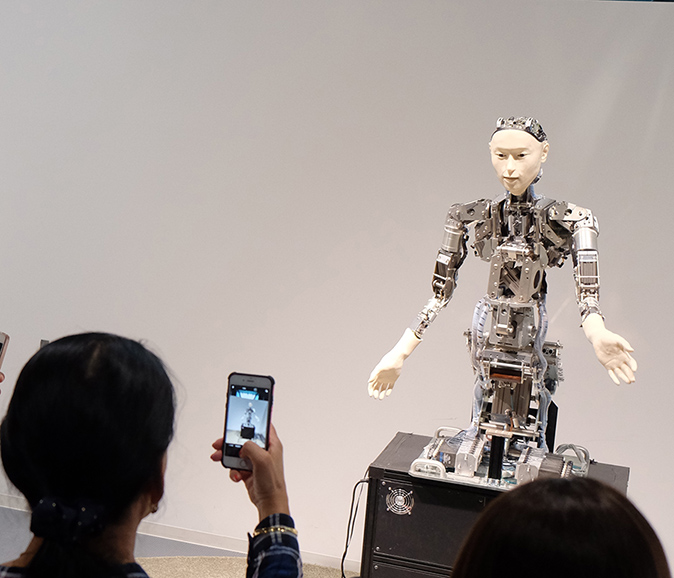

![]() The end product of development is always more complicated than the starting point. If it is not learned, it must be in the genes.
The end product of development is always more complicated than the starting point. If it is not learned, it must be in the genes.
![]() That’s not really an answer, is it? What does ‘in the genes’ really mean? I think about the genome as compressed information – 763MB for humans, in fact. The decompression requires time and energy. I think your “entire molecular code for connectivity as it develops” cannot be read in the genome.
That’s not really an answer, is it? What does ‘in the genes’ really mean? I think about the genome as compressed information – 763MB for humans, in fact. The decompression requires time and energy. I think your “entire molecular code for connectivity as it develops” cannot be read in the genome.
![]() It must be there somehow, no? If the same genome always produces a functional brain prior to any learning, where else is the information coming from?
It must be there somehow, no? If the same genome always produces a functional brain prior to any learning, where else is the information coming from?
![]() That’s what I am working on. We try to evolve AI. This is an alternative and very different approach from deep learning. If you simulate enough random genomes, you’ll end up with a network that can be smart for any specific task. But I can never predict the solutions that evolve, neither the changes in the genome nor the network. A single gene is not directly linked to the outcome, there are too many interactions and feedback. Development is like decompression – and I can’t read anything in the compressed code before decompressing it.
That’s what I am working on. We try to evolve AI. This is an alternative and very different approach from deep learning. If you simulate enough random genomes, you’ll end up with a network that can be smart for any specific task. But I can never predict the solutions that evolve, neither the changes in the genome nor the network. A single gene is not directly linked to the outcome, there are too many interactions and feedback. Development is like decompression – and I can’t read anything in the compressed code before decompressing it.
![]() Well, we are describing the decompression, I guess you could say.
Well, we are describing the decompression, I guess you could say.
![]() That’s the point! Decompression actually creates a different information content. Your genome is not a blueprint for the outcome. The genome requires time and energy to ‘grow’ new information through feedback with its own products. And I don’t think you’d be able to predict the outcome if you had no knowledge of previous outcomes.
That’s the point! Decompression actually creates a different information content. Your genome is not a blueprint for the outcome. The genome requires time and energy to ‘grow’ new information through feedback with its own products. And I don’t think you’d be able to predict the outcome if you had no knowledge of previous outcomes.
![]() I am not convinced. To me it seems it’s still in the genes. And I don’t see how your AI could become ‘human-level’ as they say without having genes or development. Bit of a big thing to leave out…
I am not convinced. To me it seems it’s still in the genes. And I don’t see how your AI could become ‘human-level’ as they say without having genes or development. Bit of a big thing to leave out…
![]() And I don’t see how AI can incorporate genes and development if all you give us are lists of molecules… I would actually argue your lists will always be incomplete, because there is always a deeper level, more interactions and more context to describe. We need to understand the decompression algorithms that generate those molecular functions! Maybe you should talk to somebody who actually tries to build something, like a robotics engineer, not just an AI researcher like me.
And I don’t see how AI can incorporate genes and development if all you give us are lists of molecules… I would actually argue your lists will always be incomplete, because there is always a deeper level, more interactions and more context to describe. We need to understand the decompression algorithms that generate those molecular functions! Maybe you should talk to somebody who actually tries to build something, like a robotics engineer, not just an AI researcher like me.
![]() We are coming from very different places. We are using different language. And I am not even sure what your problem is. I think you might need a neuroscientist to talk to as well, not only a developmental geneticist like myself.
We are coming from very different places. We are using different language. And I am not even sure what your problem is. I think you might need a neuroscientist to talk to as well, not only a developmental geneticist like myself.
![]() Well, that might not be a bad idea. There is a workshop coming up to do exactly that. We may meet both robotics engineers and neuroscientists there.
Well, that might not be a bad idea. There is a workshop coming up to do exactly that. We may meet both robotics engineers and neuroscientists there.
![]() There might be trouble ahead. What’s it called?
There might be trouble ahead. What’s it called?
![]() The Self-Assembling Brain.
The Self-Assembling Brain.
![]() Well, that’s at least what I am working on. I am not so sure about you…
Well, that’s at least what I am working on. I am not so sure about you…
![]() You may be surprised!
You may be surprised!
From the Introduction to the The Self-Assembling Brain:
“In search for answers,…
…I went to two highly respected conferences in late summer 2018, an Artificial Life conference themed “Beyond Artificial Intelligence” by the International Society for Artificial Life and the Cold Spring Harbor meeting “Molecular Mechanisms of Neuronal Connectivity.” I knew that these are two very different fields in many respects. However, my reasoning was that the artificial life and artificial intelligence communities are trying to figure out how to make something that has an existing template in biological systems. Intelligent neural networks do exist; I have seen them grow under a microscope. Surely, it must be interesting to AI researchers to see what their neurobiology colleagues are currently figuring out—shouldn’t it help to learn from the existing thing? Surely, the neurobiologists should be equally interested in seeing what AI researchers have come up with, if just to see what parts of the self-assembly process their genes and molecules are functioning in.
Alas, there was no overlap in attendance or topics. The differences in culture, language and approaches are remarkable. The neurobiological conference was all about the mechanisms that explain bits of brains as we see them, snapshots of the precision of development. A top-down and reverse engineering approach to glimpse the rules of life. By contrast, the ALifers were happy to run simulations that create anything that looked lifelike: swarming behavior, a simple process resembling some aspect of cognition or a complicated representation in an evolved system. They pursue a bottom-up approach to investigate what kind of code can give rise to life. What would it take to learn from each other? Have developmental biologists really learned nothing to inform artificial neural network design? Have Alifers and AI researchers really found nothing to help biologists understand what they are looking at? I wanted to do an experiment in which we try to learn from each other; an experiment that, if good for nothing else, would at least help to understand what it is that we are happy to ignore.
[…]
I am aware that many ALife and AI researchers may feel that reading a book written by a neurobiologist is not likely to be helpful for their work, both for reasons of perspective and the inevitable focus on unhelpful biological “messiness.” Similarly, some developmental neurobiologists may currently read a book or two on the application of deep learning to analyze their data, rather than to learn from the AI community about how “real” brains come to be. I started this project wishing there were a book that both would enjoy having a look at, or at least get sufficiently upset about to trigger discussion between the fields.